Jupyter Snippet NP ch15-code-listing
Jupyter Snippet NP ch15-code-listing
Chapter 15: Machine learning
Robert Johansson
Source code listings for Numerical Python - Scientific Computing and Data Science Applications with Numpy, SciPy and Matplotlib (ISBN 978-1-484242-45-2).
from sklearn import datasets
from sklearn import model_selection
from sklearn import linear_model
from sklearn import metrics
from sklearn import tree
from sklearn import neighbors
from sklearn import svm
from sklearn import ensemble
from sklearn import cluster
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
Built in datasets
datasets.load_boston
<function sklearn.datasets.base.load_boston(return_X_y=False)>
datasets.fetch_california_housing
<function sklearn.datasets.california_housing.fetch_california_housing(data_home=None, download_if_missing=True, return_X_y=False)>
datasets.make_regression
<function sklearn.datasets.samples_generator.make_regression(n_samples=100, n_features=100, n_informative=10, n_targets=1, bias=0.0, effective_rank=None, tail_strength=0.5, noise=0.0, shuffle=True, coef=False, random_state=None)>
Regression
np.random.seed(123)
X_all, y_all = datasets.make_regression(n_samples=50, n_features=50, n_informative=10) #, noise=2.5)
X_train, X_test, y_train, y_test = model_selection.train_test_split(X_all, y_all, train_size=0.5)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
X_train.shape, y_train.shape
((25, 50), (25,))
X_test.shape, y_test.shape
((25, 50), (25,))
model = linear_model.LinearRegression()
model.fit(X_train, y_train)
LinearRegression(copy_X=True, fit_intercept=True, n_jobs=None,
normalize=False)
def sse(resid):
return sum(resid**2)
resid_train = y_train - model.predict(X_train)
sse_train = sse(resid_train)
sse_train
1.469187171658096e-24
resid_test = y_test - model.predict(X_test)
sse_test = sse(resid_train)
sse_test
1.469187171658096e-24
model.score(X_train, y_train)
1.0
model.score(X_test, y_test)
0.31407400675201735
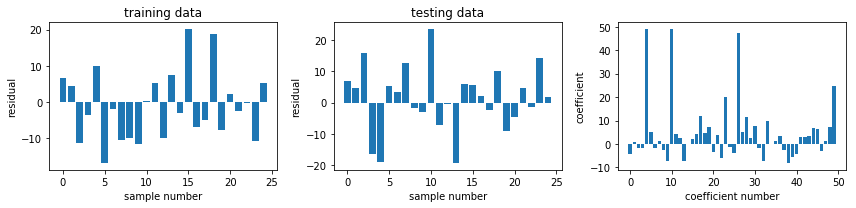
def plot_residuals_and_coeff(resid_train, resid_test, coeff):
fig, axes = plt.subplots(1, 3, figsize=(12, 3))
axes[0].bar(np.arange(len(resid_train)), resid_train)
axes[0].set_xlabel("sample number")
axes[0].set_ylabel("residual")
axes[0].set_title("training data")
axes[1].bar(np.arange(len(resid_test)), resid_test)
axes[1].set_xlabel("sample number")
axes[1].set_ylabel("residual")
axes[1].set_title("testing data")
axes[2].bar(np.arange(len(coeff)), coeff)
axes[2].set_xlabel("coefficient number")
axes[2].set_ylabel("coefficient")
fig.tight_layout()
return fig, axes
fig, ax = plot_residuals_and_coeff(resid_train, resid_test, model.coef_)
fig.savefig("ch15-regression-ols.pdf")
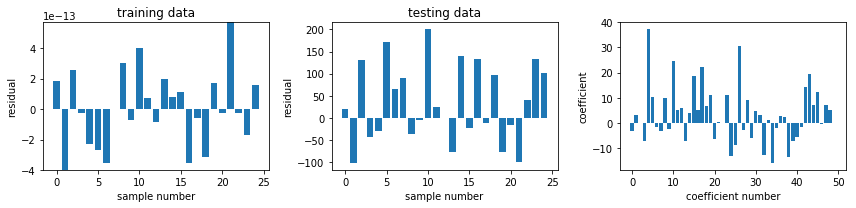
model = linear_model.Ridge() #alpha=2.5)
model.fit(X_train, y_train)
Ridge(alpha=1.0, copy_X=True, fit_intercept=True, max_iter=None,
normalize=False, random_state=None, solver='auto', tol=0.001)
resid_train = y_train - model.predict(X_train)
sse_train = sum(resid_train**2)
sse_train
178.50695164950986
resid_test = y_test - model.predict(X_test)
sse_test = sum(resid_test**2)
sse_test
212737.00160105852
model.score(X_train, y_train), model.score(X_test, y_test)
(0.9994595515017335, 0.31670332736075457)
fig, ax = plot_residuals_and_coeff(resid_train, resid_test, model.coef_)
fig.savefig("ch15-regression-ridge.pdf")
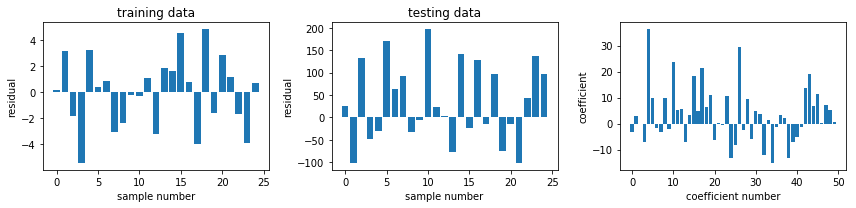
model = linear_model.Lasso(alpha=1.0)
model.fit(X_train, y_train)
Lasso(alpha=1.0, copy_X=True, fit_intercept=True, max_iter=1000,
normalize=False, positive=False, precompute=False, random_state=None,
selection='cyclic', tol=0.0001, warm_start=False)
resid_train = y_train - model.predict(X_train)
sse_train = sse(resid_train)
sse_train
309.7497138953223
resid_test = y_test - model.predict(X_test)
sse_test = sse(resid_test)
sse_test
1489.1176065002487
fig, ax = plot_residuals_and_coeff(resid_train, resid_test, model.coef_)
fig.savefig("ch15-regression-lasso.pdf")
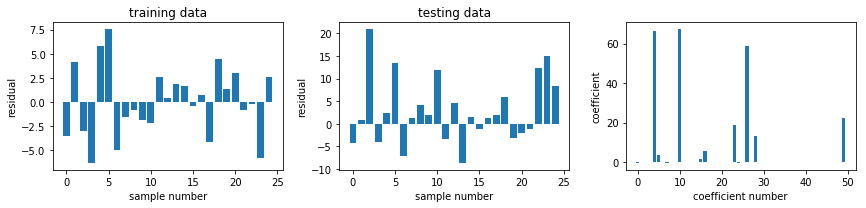
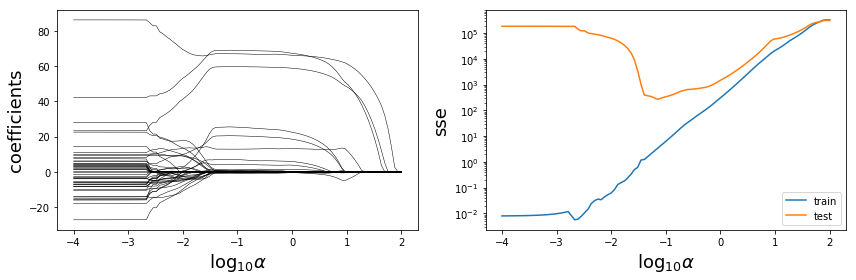
alphas = np.logspace(-4, 2, 100)
coeffs = np.zeros((len(alphas), X_train.shape[1]))
sse_train = np.zeros_like(alphas)
sse_test = np.zeros_like(alphas)
for n, alpha in enumerate(alphas):
model = linear_model.Lasso(alpha=alpha)
model.fit(X_train, y_train)
coeffs[n, :] = model.coef_
resid = y_train - model.predict(X_train)
sse_train[n] = sum(resid**2)
resid = y_test - model.predict(X_test)
sse_test[n] = sum(resid**2)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/linear_model/coordinate_descent.py:492: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations. Fitting data with very small alpha may cause precision problems.
ConvergenceWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/linear_model/coordinate_descent.py:492: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations. Fitting data with very small alpha may cause precision problems.
ConvergenceWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/linear_model/coordinate_descent.py:492: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations. Fitting data with very small alpha may cause precision problems.
ConvergenceWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/linear_model/coordinate_descent.py:492: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations. Fitting data with very small alpha may cause precision problems.
ConvergenceWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/linear_model/coordinate_descent.py:492: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations. Fitting data with very small alpha may cause precision problems.
ConvergenceWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/linear_model/coordinate_descent.py:492: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations. Fitting data with very small alpha may cause precision problems.
ConvergenceWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/linear_model/coordinate_descent.py:492: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations. Fitting data with very small alpha may cause precision problems.
ConvergenceWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/linear_model/coordinate_descent.py:492: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations. Fitting data with very small alpha may cause precision problems.
ConvergenceWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/linear_model/coordinate_descent.py:492: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations. Fitting data with very small alpha may cause precision problems.
ConvergenceWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/linear_model/coordinate_descent.py:492: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations. Fitting data with very small alpha may cause precision problems.
ConvergenceWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/linear_model/coordinate_descent.py:492: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations. Fitting data with very small alpha may cause precision problems.
ConvergenceWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/linear_model/coordinate_descent.py:492: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations. Fitting data with very small alpha may cause precision problems.
ConvergenceWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/linear_model/coordinate_descent.py:492: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations. Fitting data with very small alpha may cause precision problems.
ConvergenceWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/linear_model/coordinate_descent.py:492: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations. Fitting data with very small alpha may cause precision problems.
ConvergenceWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/linear_model/coordinate_descent.py:492: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations. Fitting data with very small alpha may cause precision problems.
ConvergenceWarning)
fig, axes = plt.subplots(1, 2, figsize=(12, 4), sharex=True)
for n in range(coeffs.shape[1]):
axes[0].plot(np.log10(alphas), coeffs[:, n], color='k', lw=0.5)
axes[1].semilogy(np.log10(alphas), sse_train, label="train")
axes[1].semilogy(np.log10(alphas), sse_test, label="test")
axes[1].legend(loc=0)
axes[0].set_xlabel(r"${\log_{10}}\alpha$", fontsize=18)
axes[0].set_ylabel(r"coefficients", fontsize=18)
axes[1].set_xlabel(r"${\log_{10}}\alpha$", fontsize=18)
axes[1].set_ylabel(r"sse", fontsize=18)
fig.tight_layout()
fig.savefig("ch15-regression-lasso-vs-alpha.pdf")

model = linear_model.LassoCV()
model.fit(X_all, y_all)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2053: FutureWarning: You should specify a value for 'cv' instead of relying on the default value. The default value will change from 3 to 5 in version 0.22.
warnings.warn(CV_WARNING, FutureWarning)
LassoCV(alphas=None, copy_X=True, cv='warn', eps=0.001, fit_intercept=True,
max_iter=1000, n_alphas=100, n_jobs=None, normalize=False,
positive=False, precompute='auto', random_state=None,
selection='cyclic', tol=0.0001, verbose=False)
model.alpha_
0.06559238747534718
resid_train = y_train - model.predict(X_train)
sse_train = sse(resid_train)
sse_train
1.5450589323146204
resid_test = y_test - model.predict(X_test)
sse_test = sse(resid_test)
sse_test
1.5321417406216717
model.score(X_train, y_train), model.score(X_test, y_test)
(0.9999953221722068, 0.9999950788657098)
fig, ax = plot_residuals_and_coeff(resid_train, resid_test, model.coef_)
fig.savefig("ch15-regression-lasso-cv.pdf")
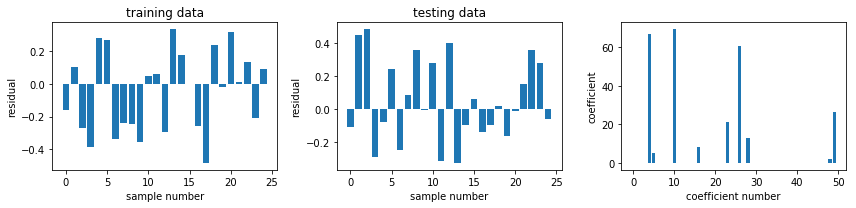
model = linear_model.ElasticNetCV()
model.fit(X_all, y_all)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2053: FutureWarning: You should specify a value for 'cv' instead of relying on the default value. The default value will change from 3 to 5 in version 0.22.
warnings.warn(CV_WARNING, FutureWarning)
ElasticNetCV(alphas=None, copy_X=True, cv='warn', eps=0.001,
fit_intercept=True, l1_ratio=0.5, max_iter=1000, n_alphas=100,
n_jobs=None, normalize=False, positive=False, precompute='auto',
random_state=None, selection='cyclic', tol=0.0001, verbose=0)
model.alpha_
0.13118477495069433
model.l1_ratio
0.5
resid_train = y_train - model.predict(X_train)
sse_train = sum(resid_train**2)
sse_train
2183.8391729391196
resid_test = y_test - model.predict(X_test)
sse_test = sum(resid_test**2)
sse_test
2650.050446338252
model.score(X_train, y_train), model.score(X_test, y_test)
(0.9933881981034111, 0.9914882195448783)
fig, ax = plot_residuals_and_coeff(resid_train, resid_test, model.coef_)
fig.savefig("ch15-regression-elastic-net-cv.pdf")
Classification
iris = datasets.load_iris()
type(iris)
sklearn.utils.Bunch
iris.target_names
array(['setosa', 'versicolor', 'virginica'], dtype='<U10')
iris.feature_names
['sepal length (cm)',
'sepal width (cm)',
'petal length (cm)',
'petal width (cm)']
iris.data.shape
(150, 4)
iris.target.shape
(150,)
# print(iris['DESCR'])
X_train, X_test, y_train, y_test = model_selection.train_test_split(iris.data, iris.target, train_size=0.7)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
classifier = linear_model.LogisticRegression()
classifier.fit(X_train, y_train)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/linear_model/logistic.py:433: FutureWarning: Default solver will be changed to 'lbfgs' in 0.22. Specify a solver to silence this warning.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/linear_model/logistic.py:460: FutureWarning: Default multi_class will be changed to 'auto' in 0.22. Specify the multi_class option to silence this warning.
"this warning.", FutureWarning)
LogisticRegression(C=1.0, class_weight=None, dual=False, fit_intercept=True,
intercept_scaling=1, max_iter=100, multi_class='warn',
n_jobs=None, penalty='l2', random_state=None, solver='warn',
tol=0.0001, verbose=0, warm_start=False)
y_test_pred = classifier.predict(X_test)
print(metrics.classification_report(y_test, y_test_pred))
precision recall f1-score support
0 1.00 1.00 1.00 16
1 1.00 0.76 0.87 17
2 0.75 1.00 0.86 12
micro avg 0.91 0.91 0.91 45
macro avg 0.92 0.92 0.91 45
weighted avg 0.93 0.91 0.91 45
np.bincount(y_test)
array([16, 17, 12])
metrics.confusion_matrix(y_test, y_test_pred)
array([[16, 0, 0],
[ 0, 13, 4],
[ 0, 0, 12]])
classifier = tree.DecisionTreeClassifier()
classifier.fit(X_train, y_train)
y_test_pred = classifier.predict(X_test)
metrics.confusion_matrix(y_test, y_test_pred)
array([[16, 0, 0],
[ 0, 12, 5],
[ 0, 0, 12]])
classifier = neighbors.KNeighborsClassifier()
classifier.fit(X_train, y_train)
y_test_pred = classifier.predict(X_test)
metrics.confusion_matrix(y_test, y_test_pred)
array([[16, 0, 0],
[ 0, 15, 2],
[ 0, 0, 12]])
classifier = svm.SVC()
classifier.fit(X_train, y_train)
y_test_pred = classifier.predict(X_test)
metrics.confusion_matrix(y_test, y_test_pred)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
array([[16, 0, 0],
[ 0, 15, 2],
[ 0, 0, 12]])
classifier = ensemble.RandomForestClassifier()
classifier.fit(X_train, y_train)
y_test_pred = classifier.predict(X_test)
metrics.confusion_matrix(y_test, y_test_pred)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
array([[16, 0, 0],
[ 0, 16, 1],
[ 0, 0, 12]])
train_size_vec = np.linspace(0.1, 0.9, 30)
classifiers = [tree.DecisionTreeClassifier,
neighbors.KNeighborsClassifier,
svm.SVC,
ensemble.RandomForestClassifier
]
cm_diags = np.zeros((3, len(train_size_vec), len(classifiers)), dtype=float)
for n, train_size in enumerate(train_size_vec):
X_train, X_test, y_train, y_test = \
model_selection.train_test_split(iris.data, iris.target, train_size=train_size)
for m, Classifier in enumerate(classifiers):
classifier = Classifier()
classifier.fit(X_train, y_train)
y_test_pred = classifier.predict(X_test)
cm_diags[:, n, m] = metrics.confusion_matrix(y_test, y_test_pred).diagonal()
cm_diags[:, n, m] /= np.bincount(y_test)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/model_selection/_split.py:2179: FutureWarning: From version 0.21, test_size will always complement train_size unless both are specified.
FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/svm/base.py:196: FutureWarning: The default value of gamma will change from 'auto' to 'scale' in version 0.22 to account better for unscaled features. Set gamma explicitly to 'auto' or 'scale' to avoid this warning.
"avoid this warning.", FutureWarning)
/Users/rob/miniconda3/envs/py3.6/lib/python3.6/site-packages/sklearn/ensemble/forest.py:246: FutureWarning: The default value of n_estimators will change from 10 in version 0.20 to 100 in 0.22.
"10 in version 0.20 to 100 in 0.22.", FutureWarning)
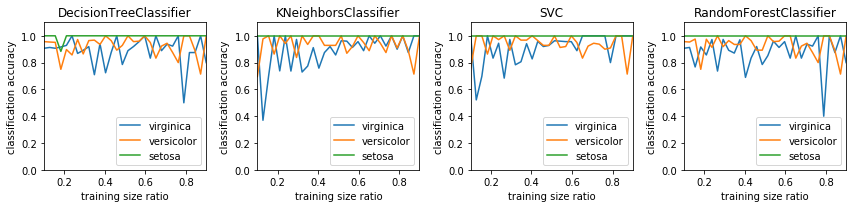
fig, axes = plt.subplots(1, len(classifiers), figsize=(12, 3))
for m, Classifier in enumerate(classifiers):
axes[m].plot(train_size_vec, cm_diags[2, :, m], label=iris.target_names[2])
axes[m].plot(train_size_vec, cm_diags[1, :, m], label=iris.target_names[1])
axes[m].plot(train_size_vec, cm_diags[0, :, m], label=iris.target_names[0])
axes[m].set_title(type(Classifier()).__name__)
axes[m].set_ylim(0, 1.1)
axes[m].set_xlim(0.1, 0.9)
axes[m].set_ylabel("classification accuracy")
axes[m].set_xlabel("training size ratio")
axes[m].legend(loc=4)
fig.tight_layout()
fig.savefig("ch15-classification-comparison.pdf")
Clustering
X, y = iris.data, iris.target
np.random.seed(123)
n_clusters = 3
c = cluster.KMeans(n_clusters=n_clusters)
c.fit(X)
KMeans(algorithm='auto', copy_x=True, init='k-means++', max_iter=300,
n_clusters=3, n_init=10, n_jobs=None, precompute_distances='auto',
random_state=None, tol=0.0001, verbose=0)
y_pred = c.predict(X)
y_pred[::8]
array([1, 1, 1, 1, 1, 1, 1, 2, 2, 2, 2, 2, 2, 0, 0, 0, 0, 0, 0],
dtype=int32)
y[::8]
array([0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 1, 1, 1, 2, 2, 2, 2, 2, 2])
idx_0, idx_1, idx_2 = (np.where(y_pred == n) for n in range(3))
y_pred[idx_0], y_pred[idx_1], y_pred[idx_2] = 2, 0, 1
y_pred[::8]
array([0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 1, 1, 1, 2, 2, 2, 2, 2, 2],
dtype=int32)
metrics.confusion_matrix(y, y_pred)
array([[50, 0, 0],
[ 0, 48, 2],
[ 0, 14, 36]])
N = X.shape[1]
fig, axes = plt.subplots(N, N, figsize=(12, 12), sharex=True, sharey=True)
colors = ["coral", "blue", "green"]
markers = ["^", "v", "o"]
for m in range(N):
for n in range(N):
for p in range(n_clusters):
mask = y_pred == p
axes[m, n].scatter(X[:, m][mask], X[:, n][mask],
marker=markers[p], s=30,
color=colors[p], alpha=0.25)
for idx in np.where(y != y_pred):
axes[m, n].scatter(X[idx, m], X[idx, n],
marker="s", s=30,
edgecolor="red",
facecolor=(1,1,1,0))
axes[N-1, m].set_xlabel(iris.feature_names[m], fontsize=16)
axes[m, 0].set_ylabel(iris.feature_names[m], fontsize=16)
fig.tight_layout()
fig.savefig("ch15-clustering.pdf")

Versions
%reload_ext version_information
%version_information sklearn, numpy, matplotlib, seaborn